Objective:
To identify the bacterial unknowns in a
mixed culture by morphological and biochemical methods.
Principle:
The process of identifying bacteria is incredibly deliberate. It involves many different methods to eliminate each a vast array of potential types – determining what, exactly is living in an unknown bacterial culture. useful benefits to many aspectofresearch, particularly the study of microorganisms and how it can be usedby physiciansto treat patient correctly. The fermentation abilities presences of certain enzymes and some biochemical reactions were determined, multiple tests were done. Observations on the test were qualitative and these tests were compared to identification keys tothe bacterial identification process based upon an unknown bacteria.
Isolation:
The importance of this step is to
isolate pure colonies of bacteria. The streak plate is a qualitative isolation
method; quadrant streaking is mostly done to obtain pure colonies. The
inoculation of the culture is made on the agar surface by back and forth
streaking with the inoculation loop over the solid agar surface. This will make
a dilution gradient across the agar plate. Upon incubation, individual colonies
will arise from the biomass.
The characteristics features of the colonies on solid agar media are:
·
Shape : circular, irregular or rhizoid.
·
Size: small, medium, large( or in
milimetres).
·
Elevation: elevated, convex, concave, umbonate/umbilicate.
·
Surface: Smooth, wavy, rough, granular,
papillate or glistening.
·
Edges: entire, undulate, crenate,
fimbriate or curled.
·
Colour: Yellow, green etc.( Note the
colour of the colony).
·
Structure: opaque, translucent or transparent.
·
Degree of growth : scanty, moderate or
profuse.
·
Nature: discrete or confluent,
filiform, spreading or rhizoid.
In order to obtain the pure culture of organism, the isolated colonies are aseptically transferred on to different nutrient agar slant tubes and incubated overnight at 37 degree Celsius. It is then stored for future purpose.
Staining Reactions:
Staining is a simple basic technique
that is used to identify microorganisms. Simple staining is used to study the
morphology of all microorganisms. The simple stain uses the basic dyes
such as Methylene blue or basic fuschin. The strong negative charge of the
bacterial cell will strongly bind with the positive charged basic dyes and will
impart its colour to all bacteria.
Fig 1: Simple staining of cocci
Gram staining is a differential staining technique that imparts different
colours to different bacteria or bacterial structures. Usually it
differentiates bacteria into two groups; gram positive and gram negative. The
primary stain Crystal violet and mordent Iodine form a strong CVI complex all
bacteria. Gram positive cells due to their thick peptidoglycan layer will
retain the CVI complex even after it is subjected to decolourization with
acetone or alcohol. Hence the counter stain Safranin has no action on gram
positive cells. But in the case of gram negative, the thin peptidoglycan layer
and more lipid contents in the cell wall will easily make them susceptible to
the action of decolorizer and hence CVI complex is easily washed out and hence
the gram negative cells will the colour of counter stain Safranin. Hence
after the gram staining, the gram positive cells appear as purple and gram negative
cells appear as pink (Fig 2). The study of morphological features and
staining characteristics help in the preliminary identification of the isolate.
Fig 2: Gram positive bacteria and Gram negative bacteria
Biochemical reactions:
Gram negative enteric bacilli play an
important role in the contamination of food. Hence they are the main causative
agents of intestinal infection. Gram negative family includes Shigella,
Salmonella, Proteus, Klebsiella, Escherichia, Enterobacter etc. Usually four
tests are used for differentiation of the various members of Enterobactericeae.
There are Indole test, Methyl red test, Voges proskauer test and Citrate test;
collectively known as IMViC series of reactions.
Indole test:
Indole tests looks for the presence or absence of tryptophanase enzyme production of the bacteria. If the enzyme is present, it will degrade the amino acid tryptophan in the media and will produce Indole, ammonia and pyruvic acid. Indole will react with Kovac's reagent to produce a cherry red complex, which indicates a positive indole test. The absence of red color is indicative of tryptophan hydrolysis due to the lack of tryptophanse enzyme(Fig 3).
Fig 3: Indole test
Methyl Red Test:
This test detects the ability of microorganism to ferment glucose and to produce acidic end products. Enteric organism produces pyruvic acid from glucose metabolism. Some enteric will then use the mixed acid pathway to metabolize pyruvic acid to other acidic products such as lactic acid, acetic acid and formic acids. This will reduce the pH of the media. Methyl red is a pH indicator which is red at the acidic pH (below 4.4) and yellow at alkaline pH (above 7). The formation of red color after the addition of Methyl red reagent indicates the accumulation of acidic end products in the medium and is an indicative of positive test (Fig 4).
Fig 4: Methyl red test
Voges Proskauer Test:
This test determines the ability of microorganism to ferment glucose. The end products of glucose metabolism,pyruvic acid, is further metabolized by using Butylene glucol pathway to produce neutral end such as acetoin and 2,3 butanediol. When Barrit's reagent A ( 40% KOH) and Barrit's reagent B (5% solution of alpha naphthol) is added it will detect the presence of acetoin, the precursor in the 2,3- butanediol synthesis. Acetoin in the presence of Oxygen and Barrit's reagent is oxidized to diacetyl, where alpha naphthol act as a catalyst. Diacetyl then reacts with guanidine components of peptone to produce a cherry red colour (Fig 5).
Citrate Utilization Test:
This test determines the ability of
microorganism to utilize Citrate. Some bacteria have the capability to convert
the salts of organic acids, for example, Sodium citrate to alkaline carbonates.
Sodium citrate is one of the important metabolite of Kreb's cycle. Certain
bacteria use citrate as the sole carbon source. Citrate utilization requires a
specific membrane transporter and citrate lyase activity. Citrate is converted
to Oxalo acetic acid by citrate lyase and oxaloacetate decarboxylase activity
will convert oxaloacetate to pyruvate with the release of carbondioxide. The
other products of the reaction are acetate, Lactic acid, formic acid etc. The
carbondioxide reacts with sodium and water to form sodium carbonate (Fig 6).
Fig 6: Citrate test
TSI:
The triple sugar- iron agar test is designed to differentiate among the different groups or genera of the Enterobacteriaceae, which are all gram negative bacilli capable of fermenting glucose with the production of acid, and to distinguish them from other gram negative intestinal bacilli. This differentiation is based on the differences in carbohydrate fermentation patterns and hydrogen sulfide production by the various groups of intestinal organisms. Carbohydrate fermentation is detected by the presence of gas and a visible colour change (from red to yellow) of the pH indicator, phenol red. The production of hydrogen sulfide is indicated by the presence of a precipitate that blackens the medium in the butt of the tube.
TSI Agar contains three fermentative sugars, lactose and sucrose in 1% concentrations and glucose in a concentration of 0.1%. Due to the building of acid during fermentation, the pH falls. The acid base indicator Phenol red is incorporated for detecting carbohydrate fermentation that is indicated by the change in colour of the medium from orange red to yellow in the presence of acids. In case of oxidative decarboxylation of peptone, alkaline products are built and the pH rises. This is indicated by the change in colour of the medium from orange red to deep red.
Sodium thiosulfate and ferrous ammonium sulfate present in the medium detects the production of hydrogen sulfide (indicated by blackening in the butt of the tube). To facilitate the detection of organisms that only ferment glucose, the glucose concentration is one-tenth the concentration of lactose or sucrose. The small amount of acid produced in the slant of the tube during glucose fermentation oxidizes rapidly, causing the medium to remain orange red or revert to an alkaline pH.
In contrast, the acid reaction (yellow) is maintained
in the butt of the tube since it is under lower oxygen tension. After depletion
of the limited glucose, organisms able to do so will begin to utilize the
lactose or sucrose. To enhance the alkaline condition of the slant, free
exchange of air must be permitted by closing the tube cap loosely. If the tube
is tightly closed, an acid reaction (caused solely by glucose fermentation)
will also involve the slant.
Urease test:
Urea is a nitrogen containing compound
that is produced during the decarboxylation process of the amino acid arginine
in the urea cycle. Urea is highly soluble in water and is thus it is an
efficient way for the human body to excess nitrogen. This excess urea is then
taken away from the body with the help of the kidneys as a part of urine.
Certain bacteria produce the enzyme urease during its metabolism process and
that will break down the urea in the medium to ammonia and carbon dioxide:
Some enteric bacteria produce the
enzyme urease, which splits the urea molecule into carbon dioxide and ammonia.
The urease test is useful in identifying the genera Proteus, Providentia, and
Morganella, which liberate this enzyme.
Urease is a hydrolytic enzyme that
attacks the nitrogen and carbon bond in amide compounds such as urea and forms
the alkaline end product ammonia. The presence of urease is detectable when the
organisms are grown in a urea broth medium containing the pH indicator phenol
red. As the substrate urea is split into its products, the presence of ammonia
creates an alkaline environment that causes the phenol red to turn to deep
pink. This is a positive reaction for the presence of urease. Failure of deep
pink colour to develop is evidence of a negative reaction.
SIM:
The ability of an organism to move by itself is called motility. Motility is closely linked with chemotaxis, the ability to orientate along certain chemical gradients. Eucaryotic cells can move by means of different locomotor organelles such as cilia, flagella, or pseudopods. Prokaryotes move by means of propeller-like flagella unique to bacteria or by special fibrils that produce a gliding form of motility. Almost all spiral bacteria and about half of the bacilli are motile, whereas essentially none of the cocci are motile.
The medium mainly used for this purpose is SIM medium (Sulphide Indole Motility medium) which is a combination differential medium that tests three different parameters, sulphur reduction, indole production and motility. This media has a very soft consistency that allows motile bacteria to migrate readily through them causing cloudiness. In soft agar tubes non-motile bacteria will only grow on the inoculated region. Motile bacteria will grow along the stab line and will tend to swim out away from the stabbed area. Therefore, a negative result is represented by growth in a distinct zone directly along the stab. A positive result is indicated by diffuse or cloudy growth mainly at the top and bottom of the stabbed region.
SIM agar may also be used to detect the
presence of H2S production. The SIM medium contains peptones
and sodium thiosulfate as substrates, and ferrous
ammonium sulfate, Fe(NH4)SO4, as the H2S
indicator. Cysteine is a component of the peptones used in SIM medium.
Sufficient agar is present to make the medium semisolid. Once
H2S is produced, it combines with the
ferrous ammonium sulfate, forming an insoluble, black ferrous sulfide
precipitate that can be seen along the line of the stab inoculation. If the
organism is also motile, the entire tube may turn black. This black
line or tube indicates a positive H2S reaction; absence of a
black precipitate indicates a negative reaction.
Gelatin Hydrolysis Test:
Gelatin, a protein derived from the animal
protein collagen. It has been used as a solidifying agent in food for a long
time. Robert Koch used nutrient gelatin as an early type of solid growth
medium. One problem is that many bacteria have the ability to hydrolyze
(liquefy) gelatin. This gelatin liquefaction ability (or inability) forms the
basis for this test. Some microorganisms possess an enzyme called gelatinase,
which breaks down gelatin into amino acids. Gelatin deeps contain the substrate
gelatin, which is a protein produced by the hydrolysis of collagen. Organisms
which hydrolyze gelatin will cause the gelatin to liquefy.
The gelatin hydrolysis tests for an organism's ability to break down the protein gelatin which is derived from collagen. Gelatin causes the media to thicken, especially at cooler (below 28oC) temperatures. If the organism can release gelatinase enzymes the gelatin is broken down or liquefied. The media is checked over a period of about a week after inoculation and incubation at room temperature, for gelatinase activity. The tube is placed on ice for a few minutes and if the media fails to solidify it is considered a positive test. The gelatinase reaction may be slow or incomplete (Fig 7).
Fig 7: Gelatin hydrolysis test (The gelatin hydrolysis is indicated by the liquid nature of the gelatin ( positive test) and the negative test is indicated by the solid nature of the gelatin)
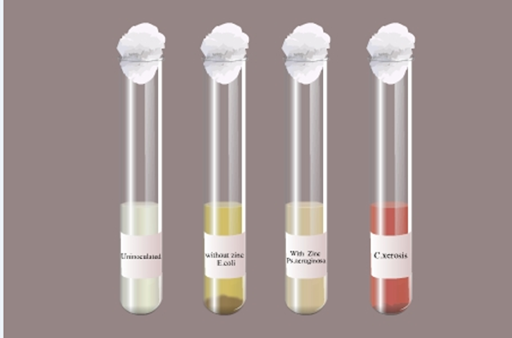
Nitrate Reduction Broth:
Bacterial species may be classified
into different groups depends on their ability to reduce nitrate to
nitrite or nitrogenous gases provided in the growth medium. The reduction in
nitrate can be coupled to anaerobic respiration in some bacterial
species. Nitrate, present in the broth, is reduced to nitrite which
is then reduced to nitric oxide, nitrous oxide, or nitrogen. The basis of
nitrate reduction test the detection of nitrite and its ability to form a red
colored compound when it reacts with reagent A which is sulfanilic
acid and to form a complex (nitrite-sulfanilic acid) which then reacts
with Reagent B which is α-naphthylamine to give a red precipitate (prontosil).
Zinc powder act as a catalyst and that will favours the reduction of nitrate to
nitrite. Nitrate reaction occurs only under anaerobic conditions (Fig 8). The
medium is then transferred in tubes to make a low surface area to depth ratio
that will limit the diffusion of oxygen into the growth medium. Most bacteria
utilize the available oxygen in the medium for their growth and will rapidly
produce anaerobic conditions for the further reactions.
Fig 8: Nitrate reduction test
Catalase Test:
The inability of strict anaerobes to
synthesize catalase, peroxidase, or superoxide dismutase may explain why oxygen
is poisonous to these microorganisms. In the absence of these enzymes, the
toxic concentration of H2O2 cannot be degraded when
these organisms are cultivated in the presence of oxygen. Organisms capable of
producing catalase rapidly degrade hydrogen peroxide which is a tetramer
containing four polypeptide chains, which are usually 500 amino acids
long. It also contains four porphyrin heme groups(ie., iron groups) that will
allow the enzyme to react with the hydrogen peroxide.
The enzyme catalase is present in most
cytochrome containing aerobic and facultative anaerobic bacteria. Catalase has
one of the highest turnover numbers of all enzymes such that one molecule of
catalase can convert millions of molecules of hydrogen peroxide to water and
oxygen in a second.
Catalase production and activity can be
detected by adding the substrate H2O2 to an
appropriately incubated (18- to 24-hour) tryptic soy agar slant culture.
Organisms which produce the enzyme break down the hydrogen peroxide, and the
resulting O2 production produces bubbles in the reagent drop,
indicating a positive test. Organisms lacking the cytochrome system also lack
the catalase enzyme and are unable to break down hydrogen peroxide, into O2 and
water and are catalase negative.
Coagulase Test:
Coagulases are enzymes that clot blood
plasma by a mechanism that is similar to normal clotting. The coagulase test
identifies whether an organism produces this exoenzyme. This enzyme clots the
plasma component of blood. The only significant disease-causing bacteria of
humans that produce coagulase are Staphylococcus aureus. Thus this
enzyme is a good indicator of the pathogenic potential of S. aureus. In the
test, the sample is added to rabbit plasma and held at 37° C for a specified
period of time. Formation of clot within 4 hours is indicated as a positive
result and indicative of a virulent Staphylococcus aureus strain.
The absence of coagulation after 24 hours of incubation is a negative result,
indicative of an avirulent strain.
Oxidase Test:
Oxidase test is an important differential procedure that should be performed on
all gram-negative bacteria for their rapid identification. The test depends on
the ability of certain bacteria to produce indophenol blue from the oxidation
of dimethyl-p-phenylenediamine and α-naphthol. This method uses N,N-dimethyl-p-phenylenediamine
oxalate in which all Staphylococci were oxidase negative. In presence of the
enzyme cytochrome oxidase (gram-negative bacteria) the
N,N-dimethyl-p-phenylenediamine oxalate and α-naphthol react to indophenol
blue. Pseudomonas aeruginosa is an oxidase positive organism.
Starch Hydrolysis Test:
Amylases are a class of enzymes that are capable of digesting these glycosidic
linkages found in starches. Amylases can be derived from a variety of sources.
Amylases are present in all living organisms, but the enzymes vary in activity,
specificity and requirements from species to species and even from tissue to
tissue in the same organism. Alpha-amylase (1,4 alpha
D-Glucan-glucanohydrolase) acts upon large polymers of starch at internal bonds
and cleaves them to short glucose polymers. Alpha-amylase catalyzes the
hydrolysis of internal Alpha-1-4 glucan bonds in polysaccharides containing 3
or more alpha 1-4 linkages; it results in a mixture of maltose and glucose.
Amyloglucosidase works on the shorter polymers and splits off single glucose
sugars. Bacterial alpha-amylase is particularly suited for industrial usage
since it is inexpensive and isothermally stable.
Starch agar is an example of differential medium which tests the ability of an
organism to produce certain alpha-amylase and oligo-1, 6-glucosidase that
hydrolyze starch. Starch molecules are too large to enter into the bacterial
cells, so some bacteria will secrete exoenzymes that will degrade starch into
subunits that can be then easily utilized by the organism.
Starch agar is a simple nutritive medium with starch added. Since no
colour change occurs in the medium when organisms hydrolyze starch, iodine
solution is added to the plate after incubation. Iodine turns blue,
purple, or black (the colour depends on the concentration of the iodine used)
in the presence of starch. A clearing around the bacterial growth shows that
the organism has hydrolyzed starch.



















0 Comments